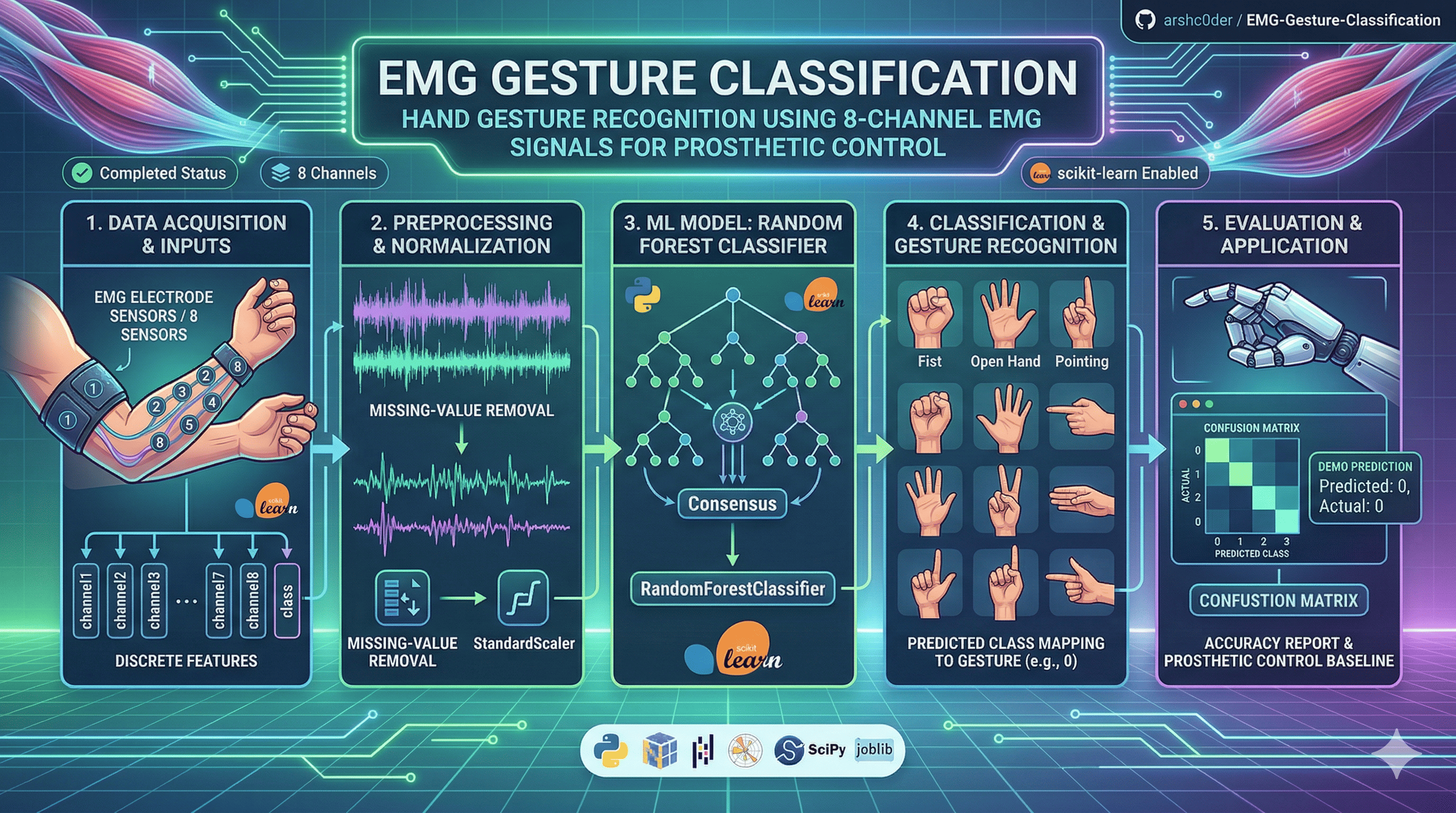
This project builds a machine learning pipeline for EMG signal classification using 8-channel electromyography (EMG) data.
The goal is to classify hand gesture classes for prosthetic-control style applications, where muscle activity signals are used to recognize intended movement patterns.
This project demonstrates a practical biosignal classification workflow including:
- dataset loading
- preprocessing
- normalization
- model training
- evaluation
- confusion matrix generation
- demo prediction
The main goals of this project are to:
- classify hand gesture classes from EMG signals
- build a baseline prosthetic-control style classifier
- demonstrate biosignal preprocessing and machine learning workflow
- generate clear evaluation outputs for model performance analysis
The model uses:
- Input features:
channel1tochannel8 - Target label:
class - Model:
RandomForestClassifier - Preprocessing: missing-value removal + normalization using
StandardScaler
This makes the project a strong starting point for EMG-based gesture recognition systems.
Predicted class: 0
Actual class: 0
Input values: {
'channel1': -2e-05,
'channel2': 1e-05,
'channel3': 0.0,
'channel4': -7e-05,
'channel5': -3e-05,
'channel6': 1e-05,
'channel7': 0.0,
'channel8': -1e-05
}
This example shows that the trained model correctly predicted the class for one test sample.
The dataset file is stored in compressed form to keep the repository lightweight.
After extraction, the project should contain:
data/EMG-data.csvYou can keep the dataset in the repository as:
data/EMG-data.zipor
data/EMG-data.csv.zipThen extract it so the CSV becomes:
data/EMG-data.csvThe raw CSV file can be relatively large, while the zip file is much smaller.
Keeping it compressed helps to:
- reduce repository size
- improve cloning and downloading speed
- make the project easier to share with contributors
EMG-Gesture-Classification/
│
├── data/
│ ├── EMG-data.zip # or EMG-data.csv.zip
│ └── EMG-data.csv # extracted dataset file
│
├── models/
│ ├── emg_random_forest.pkl
│ └── scaler.pkl
│
├── results/
│ ├── accuracy_report.txt
│ ├── confusion_matrix.png
│ └── demo_prediction.txt
│
├── train_emg.ipynb
├── README.md
└── .gitignore
The classification workflow includes:
- loading EMG data
- selecting channels
channel1tochannel8 - using
classas the target label - removing missing rows
- normalizing feature values
- splitting data into training and testing sets
- training a Random Forest model
- evaluating predictions
- saving model and result files
- Python
- NumPy
- Pandas
- Matplotlib
- scikit-learn
- SciPy
- joblib
Install Python 3.10+ and required libraries:
pip install numpy pandas matplotlib scikit-learn scipy joblibpython -m venv emg_env
.\emg_env\Scripts\Activate.ps1
pip install numpy pandas matplotlib scikit-learn scipy joblibpython -m venv emg_env
emg_env\Scripts\activate.bat
pip install numpy pandas matplotlib scikit-learn scipy joblibpython3 -m venv emg_env
source emg_env/bin/activate
pip install numpy pandas matplotlib scikit-learn scipy joblibExpand-Archive -Path .\data\EMG-data.zip -DestinationPath .\data -Forceunzip data/EMG-data.zip -d dataExpand-Archive -Path .\data\EMG-data.csv.zip -DestinationPath .\data -Forceunzip data/EMG-data.csv.zip -d dataAfter extraction, verify that this file exists:
data/EMG-data.csvIf you are using the notebook:
jupyter notebookThen open:
train_emg.ipynband run all cells.
Since this project uses a notebook, avoid writing
python train_emg.ipynb. If you later convert it to a script, you can use:python train_emg.py
After running the project, the following files are generated:
results/
├── accuracy_report.txt
├── confusion_matrix.png
└── demo_prediction.txt
models/
├── emg_random_forest.pkl
└── scaler.pkl
Contains:
- overall model accuracy
- classification report
- class-wise performance summary
Shows how well the model predicts each class.
Shows one sample prediction from the test set.
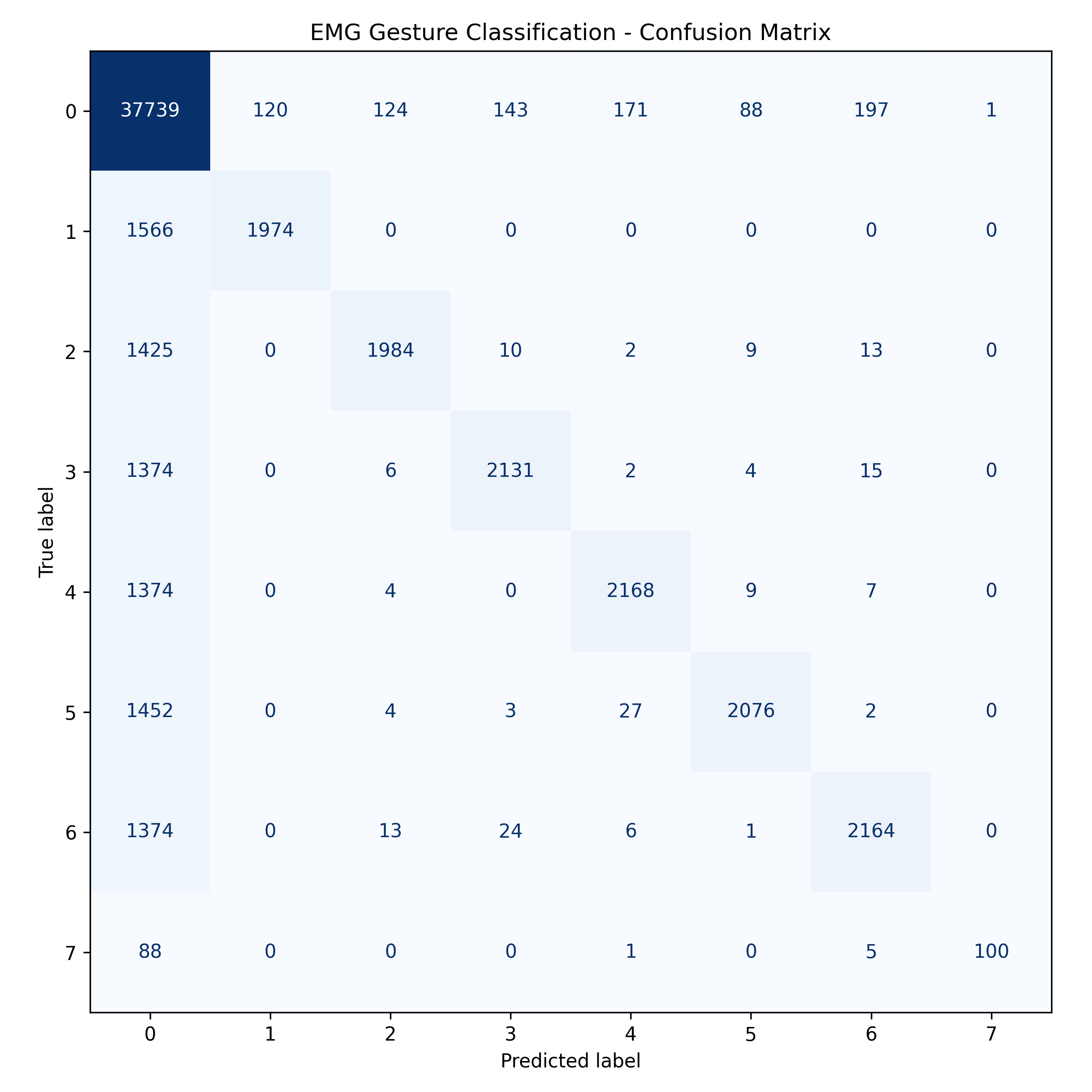
This matrix helps visualize how well the classifier distinguishes between gesture classes.
The baseline model uses these EMG channels as input:
channel1channel2channel3channel4channel5channel6channel7channel8
class
- This baseline version predicts the numeric values stored in the
classcolumn. - The
timeandlabelcolumns are not used in the first version. - If a mapping between class IDs and real gesture names becomes available later, the project can be upgraded to display gesture names instead of numeric classes.
- EMG biosignal preprocessing
- multi-channel feature-based classification
- normalization for stable model training
- supervised machine learning workflow
- confusion matrix based evaluation
- prosthetic-control style gesture recognition baseline
- clean baseline machine learning pipeline
- practical biosignal classification use case
- easy-to-understand notebook workflow
- reusable saved model and scaler
- strong base for future signal-processing improvements
-
add EMG signal filtering
-
implement window-based segmentation
-
extract engineered EMG features such as:
- MAV
- RMS
- Variance
- Waveform Length
-
compare with SVM, KNN, and XGBoost
-
map numeric predictions to gesture names
-
add live prediction support
-
build a prosthetic-control demo interface
-
evaluate feature importance across EMG channels
This kind of EMG gesture classification system can be extended for:
- prosthetic hand control
- human-computer interaction
- wearable biosignal interfaces
- rehabilitation systems
- gesture-based assistive technology
__pycache__/
*.pyc
emg_env/
models/
results/
data/*.csvAdjust this depending on whether you want model files or images pushed to GitHub.
Contributions are welcome.
If you'd like to improve preprocessing, add feature extraction, compare classifiers, or build a real-time EMG application, feel free to fork the repository and open a pull request.
This project is licensed under the MIT License.
arshc0der
This project is a practical implementation of:
- EMG signal classification
- hand gesture recognition
- prosthetic-control style AI
- biosignal preprocessing
- machine learning classification
- performance evaluation with confusion matrix
If you found this project useful, consider giving it a star on GitHub.