IMPERANDI is a Python framework and CLI for building analysis-ready CT imaging cohorts from heterogeneous DICOM sources. It standardizes identifiers, curates volume-level metadata, converts volumes to NIfTI, and supports downstream segmentation, perfusion phase detection, radiomics extraction, and quality control in one coherent pipeline.
This work performed under the RHU OPERANDI project was supported in part by the French National Research Agency (Agence Nationale de la Recherche, ANR) as its 3rd PIA, integrated to France 2030 plan under reference ANR-21-RHUS-0012.
- Reduces manual data wrangling by turning raw DICOM trees into structured cohort tables.
- Improves reproducibility with explicit CSV outputs at every stage and deterministic ID logic.
- Improves reliability on real hospital exports with archive support, failure tracking, and resumable workflows.
- Keeps adoption practical in secure environments with a lightweight Python-first toolchain.
- Scans DICOM files from folders, globbed roots, and nested archives (
.zip,.tar,.tar.gz,.tgz). - Extracts selected DICOM header tags into a raw metadata table (
dicom_index.csv). - Builds stable patient/study/series identifiers from tags, folder structure, or hybrid fallback rules.
- Applies manifest-driven hooks for patient-key standardization and derived columns.
- Cleans and curates CT cohorts by filtering modality/noise patterns, localizers, non-target anatomy, non-axial acquisitions, and implausible scan geometry.
- Aggregates slices into robust volume-level records and computes exam/acquisition ordering.
Impact: turns fragmented acquisition data into a consistent cohort backbone that downstream models and analytics can trust.
- Converts curated DICOM volume rows to NIfTI in parallel using
dicom2nifti. - Preserves source-to-output traceability in a CSV (
nifti_pathper row). - Handles archive-backed DICOM paths transparently via on-demand materialization.
- Writes explicit conversion error tables without aborting the whole run.
Impact: creates a standardized imaging representation for model training, segmentation, and feature extraction at scale.
- Runs configurable task pipelines (default backend: TotalSegmentator).
- Supports multi-task mask generation per volume through a JSON task config.
- Adds optional post-processing (mask merge, closing, hole filling, largest connected component).
- Uses multiprocessing with timeout controls and produces warning/error tracking CSVs.
Impact: converts raw CT volumes into ready-to-use anatomical/tumor masks with operational safeguards for large cohort processing.
- Extracts CT contrast phase metadata from NIfTI volumes using TotalSegmentator phase utilities.
- Appends normalized phase outputs to cohort CSVs (
totalseg_*columns). - Captures per-row failures into dedicated error outputs.
Impact: enables phase-aware stratification and analysis without manual review of every study.
- Extracts PyRadiomics features for organ and tumor regions from CT + masks.
- Includes a organ-minus-tumor extraction path for cleaner parenchyma characterization.
- Supports optional cohort filtering controls and error-aware output generation.
- Supports PyRadiomics parameterization from either
--pyradiomics_settings /path/to/Params.yamlor manifestradiomicssettings.
Impact: accelerates feature exctraction for prognostic and response modeling pipelines.
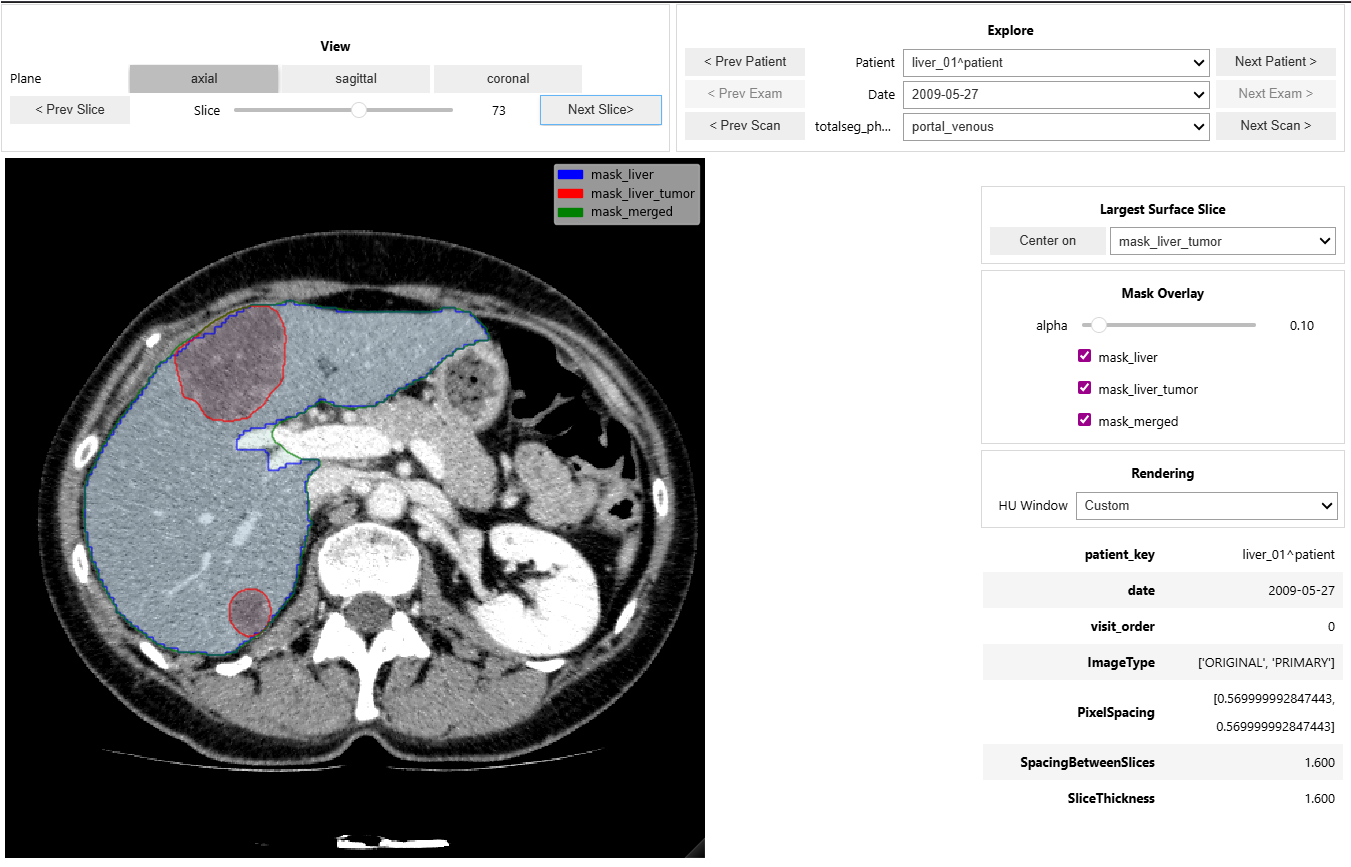
- Provides an interactive CT + mask viewer for cohort navigation and quick visual QA.
- Supports patient/date/phase exploration, mask overlays, window presets, and keyboard navigation.
Impact: shortens the feedback loop between pipeline outputs and clinical/imaging validation.
IMPERANDI ships a single CLI with these subcommands:
parse: scan DICOMs and build metadata index tables.clean: filter and normalize parsed metadata.ingest: runparsethenclean.convert: convert indexed DICOM volumes to NIfTI.segment: run configurable segmentation on NIfTI volumes (requires TotalSegmentator, install with.[segment]).phase: extract contrast phase metadata from NIfTI volumes (requires TotalSegmentator, install with.[segment]).radiomics: extract radiomics features from NIfTI volumes and masks (requires pyRadiomics, install with.[radiomics]).
Get help:
imperandi --help
imperandi parse --help
imperandi clean --help
imperandi ingest --help
imperandi convert --help
imperandi segment --help
imperandi phase --help
imperandi radiomics --helpBase install:
python -m pip install -e .Segmentation dependencies:
python -m pip install -e ".[segment]"Radiomics dependencies:
python -m pip install -e ".[radiomics]"Development and test tooling:
python -m pip install -e ".[dev]"Enable tracked git hooks (recommended):
git config core.hooksPath .githooksWith hooks enabled, git push strips output/execution state from changed *.ipynb files, stages those changes, and stops once so you can commit the cleaned notebooks.
Install everything:
python -m pip install -e ".[all]"Optional Jupyter kernel setup:
python -m ipykernel install --user --name imperandi310 --display-name "IMPERANDI (Python 3.10)"Run ingest (parse + clean):
imperandi ingest \
--root_path /path/to/dicom \
--output_dir /path/to/output \
--manifest genericConvert to NIfTI:
imperandi convert \
--csv_path /path/to/output/dicom_index_clean.csv \
--output_dir /path/to/nifti_root \
--csv_path_out /path/to/output/nifti_index.csvRun segmentation:
imperandi segment \
--csv_path /path/to/output/nifti_index.csv \
--csv_path_out /path/to/output/nifti_index_segmented.csvExtract contrast phase:
imperandi phase \
--csv_path /path/to/output/nifti_index_segmented.csv \
--csv_path_out /path/to/output/nifti_index_phased.csvExtract radiomics:
imperandi radiomics \
--csv_path /path/to/output/nifti_index_segmented.csv \
--csv_path_out /path/to/output/nifti_index_radiomics.csvExtract radiomics with explicit PyRadiomics YAML settings:
imperandi radiomics \
--csv_path /path/to/output/nifti_index_segmented.csv \
--pyradiomics_settings /path/to/Params.yaml \
--csv_path_out /path/to/output/nifti_index_radiomics.csvUse manifest-defined radiomics settings:
imperandi radiomics \
--csv_path /path/to/output/nifti_index_segmented.csv \
--manifest generic \
--csv_path_out /path/to/output/nifti_index_radiomics.csvIf both --manifest and --pyradiomics_settings are provided, IMPERANDI warns and
prefers manifest radiomics settings when that section exists.
parse:dicom_index.csv(resolved IDs and selected DICOM tags)- optional
dicom_tags_snapshot.ndjson(full recursive tags on a sampled subset, via--snapshot_tags)
clean:- cleaned cohort table (default
<input>_clean.csv)
- cleaned cohort table (default
convert:- NIfTI-enriched cohort table (
nifti_index.csvby default) - conversion failures (
conv_errors.csvby default)
- NIfTI-enriched cohort table (
segment,phase,radiomics:- enriched cohort table + command-specific error CSV
Manifests define dataset-specific behavior and live in:
src/imperandi/datasets_config/manifests/*.json
Hook implementations live in:
src/imperandi/datasets_config/hooks/
You can pass either a manifest name (generic, operandi) or a custom manifest path.
For radiomics, manifest key radiomics can directly contain a PyRadiomics-style
settings object (same structure as Params.yaml content).
- Parallel execution controls are available for heavy stages (
parse,convert,segment). - Long-running stages (
parse,convert,segment,phase,radiomics) use a unified checkpoint interface:--checkpoint_every_rows,--checkpoint_every_sec,--no_resume,--strict_resume. - Resume is enabled by default; pass
--no_resumeto disable it. parsereads tags from defaults (DEFAULT_DICOM_TAGS) plus--tags; use--snapshot_tagsfor full recursive tag snapshots on sampled data.parseauto-detects archive-heavy inputs from a deterministic root sample (--archive_detect_sample_size) and can switch to archive-aware mode at runtime when needed.- Archive workflows are bounded by depth and include path-safety protections.
- Most commands support
--dry-runfor pipeline planning and CI smoke checks.
Use Case on IRCAD Dataset
Download the dataset (~800MB):
wget https://cloud.ircad.fr/index.php/s/JN3z7EynBiwYyjy/download -O ircad.zipUnzip the archive:
unzip ircad.zip -d ircad_dicomAfter extraction, your structure should look similar to:
ircad_dicom/
└── 3Dircadb1/
├── 3Dircadb1.1/
│ ├── PATIENT_DICOM.zip/
│ ├── MASKS_DICOM.zip/
│ └── ...
Install package:
conda create -n imperandi310 python=3.10
conda activate imperandi310
pip install -e .[all]Execute pipeline:
imperandi ingest "ircad_dicom/3Dircadb1/**/PATIENT_DICOM*" . --snapshot_tags
imperandi convert dicom_index_clean.csv ircad_nifti/
imperandi segment nifti_index.csv
imperandi phase nifti_index.csv
imperandi radiomics nifti_index.csvInspect results with dashboards:
- explore images & segmentations with the interactive viewer
- inspect DICOM tags
- basic radiomics statistics